 
In brief, it addresses the challenges identified above in five major steps: (a) orthogroup inference, (b) inference of gene trees for each orthogroup, (c and d) analysis of these gene trees to infer the rooted species tree, (e) rooting of the gene trees. For doing so, it compares similarities of given gene sequences and clusters them to. The OrthoFinder algorithm is described in detail in the Methods section. Proteinortho is a tool to detect orthologous genes within different species. 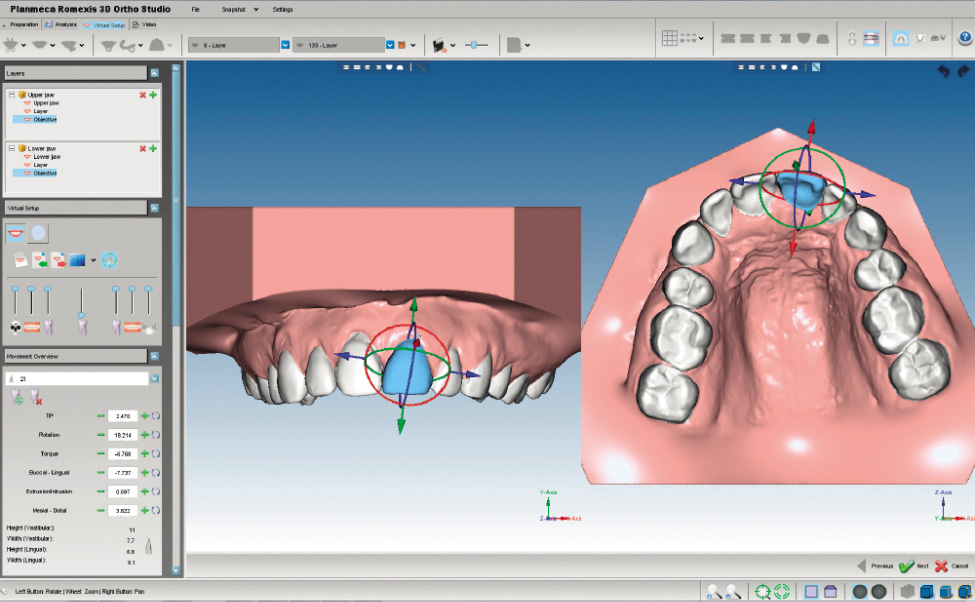
These gene regions may have played critical roles in the adaptation of seagrasses to diverse environments. 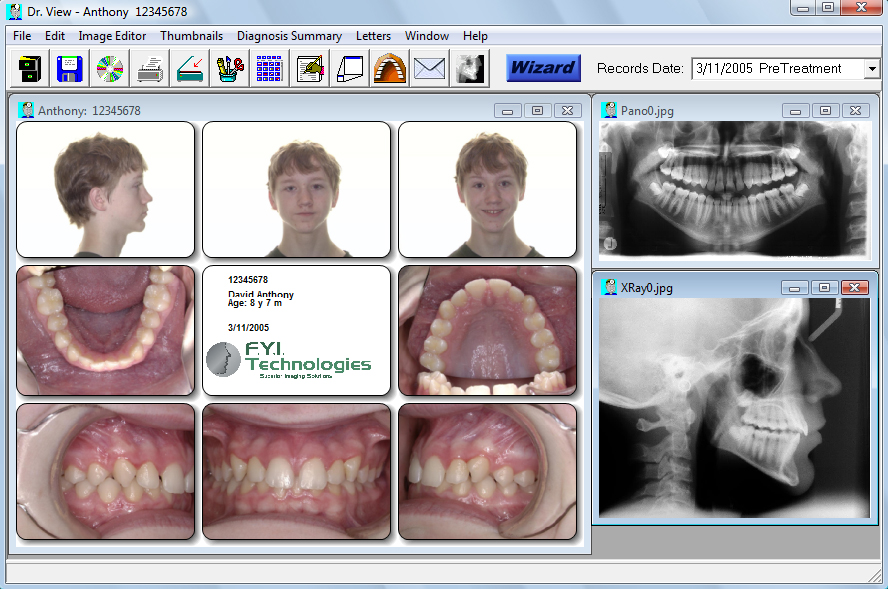
SAGE and EST library data, such as the Cancer Genome. Remotes, knitr, BiocStyle, covr, markdown, rmarkdown, here, testthat (>= 3.0.0), piggyback, badger, magick, desc, hrbrthemes, Cairo, yulab. OrthoFinder algorithm overview and summary of results files. Interestingly, we found nine genes with positive selection sites, including two ATP subunit genes (atpA and atpF), three ribosome subunit genes (rps4, rps7, and rpl20), one photosystem subunit gene (psbH), and the ycf2, accD, and rbcL genes. Alternatively, one could extract this information from publicly available. To extract the DNA you will use a detergent called sodium dodecyl sulfate (SDS) to remove the. 
To view documentation for the version of this package installedĬomparativeGenomics, GeneExpression, Genetics, Phylogenetics, Preprocessing, Software, Transcriptomicsĭplyr, methods, stats, utils, Matrix, jsonlite, homologene, gprofiler2, babelgene, data.table, parallel, ggplot2, ggpubr, patchwork, DelayedArray, grr, repmis, ggtree, tools Explain the relationship among DNA, genes and chromosomes. If (!require("BiocManager", quietly = TRUE))įor older versions of R, please refer to the appropriate To install this package, start R (version 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
